Interpretable Explainable machine learning for
time-to-event prediction in medicine and healthcare
Hubert Baniecki
University of Warsaw, Poland
BIRS Workshop, Banff, Canada
February 12, 2024
👋 Hi!
PhD student at the University of Warsaw, Poland
explainable machine learning, evaluating explanations, robustness
statistical software: dalex (JMLR 2021, John M. Chambers Award by ASA), modelStudio (JOSS 2019), survex (Bioinformatics 2023)


📣 How to explain machine learning survival models?
- method: time-dependent explanations (Knowledge-Based Systems 2023)
- software: an R package
survex(Bioinformatics 2023) - application(s): finding bias in predictions from medical data (AIME 2023)




🔨 method:
time-dependent explanations
How to explain machine learning survival models?
For classification (or regression), we have:
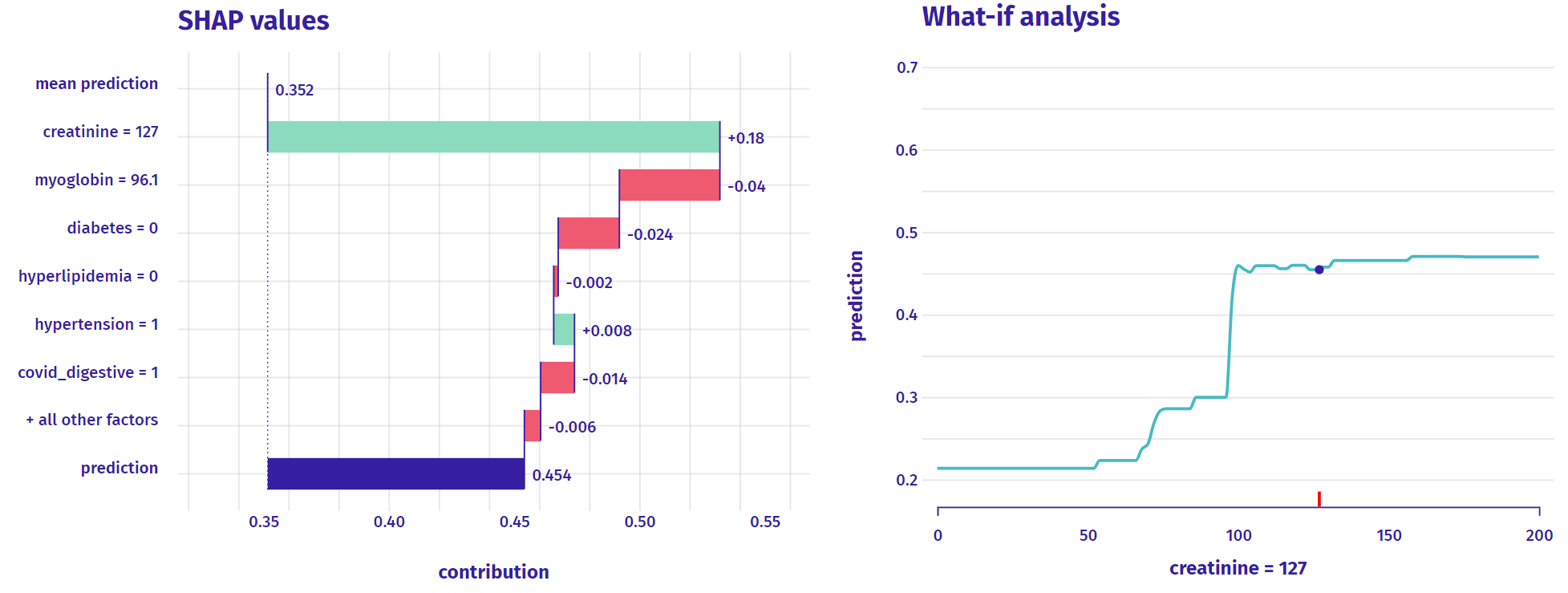
What about time-to-event prediction like survival analysis?
SurvLIME
(Ribeiro et al. 2016) LIME idea: approximate a black-box function with an interpretable model in the local neighbourhood of observation \(x\).
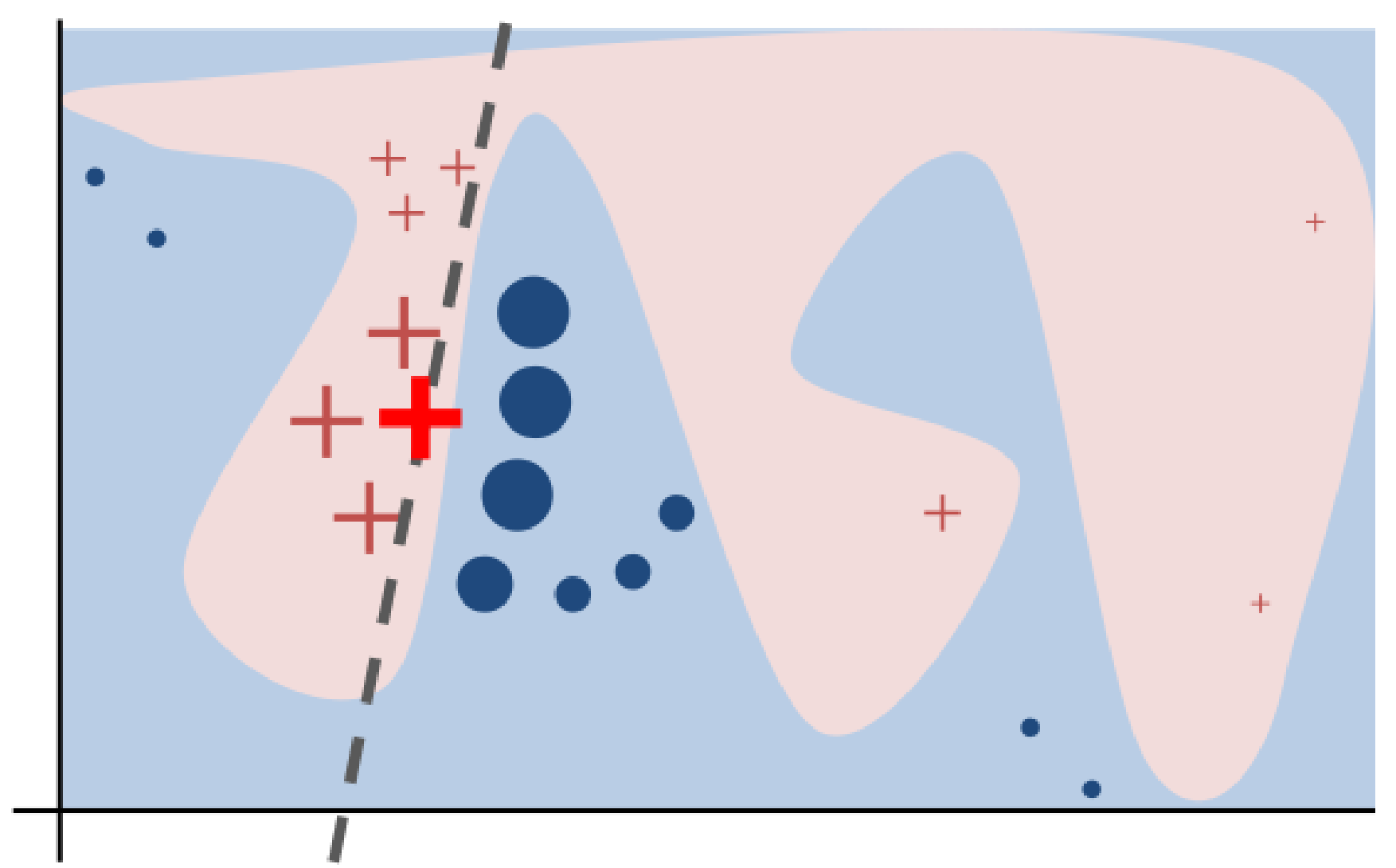

M. S. Kovalev et al. SurvLIME: A method for explaining machine learning survival models. Knowledge-Based Systems 2020
SurvLIME: example and limitations
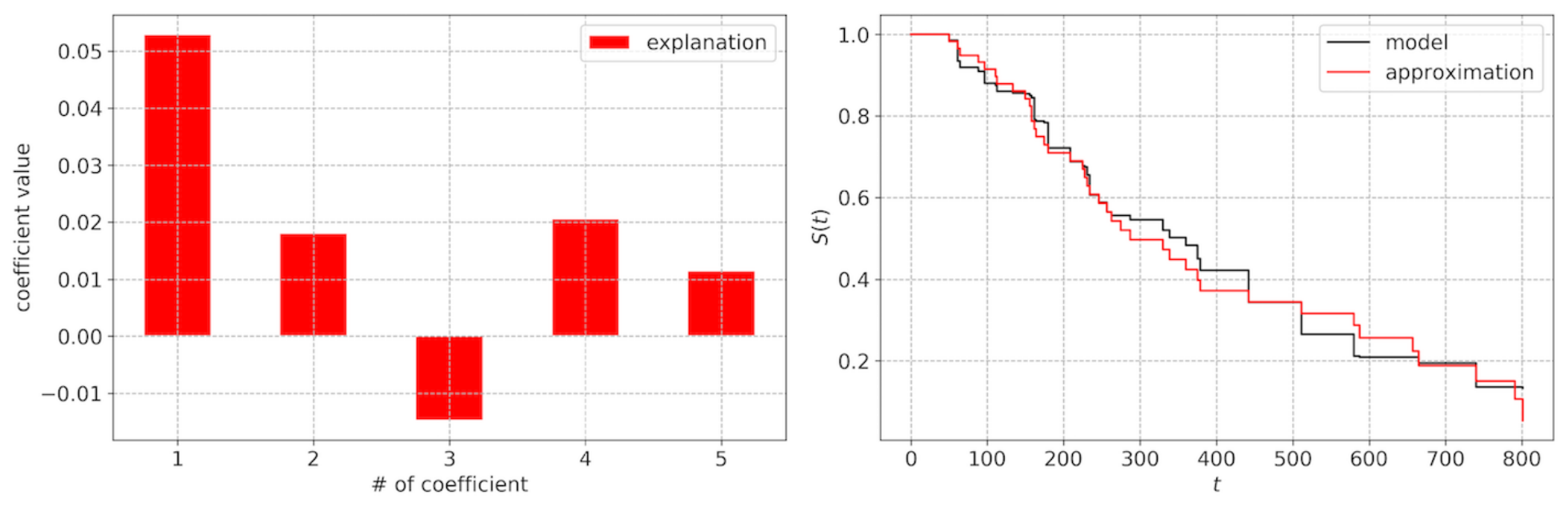
- an explanation lacks the time dimension
- approximating a complex model with linear coefficients (local accuracy)
- coefficients are not importances (depend on feature values)
SurvSHAP(t)
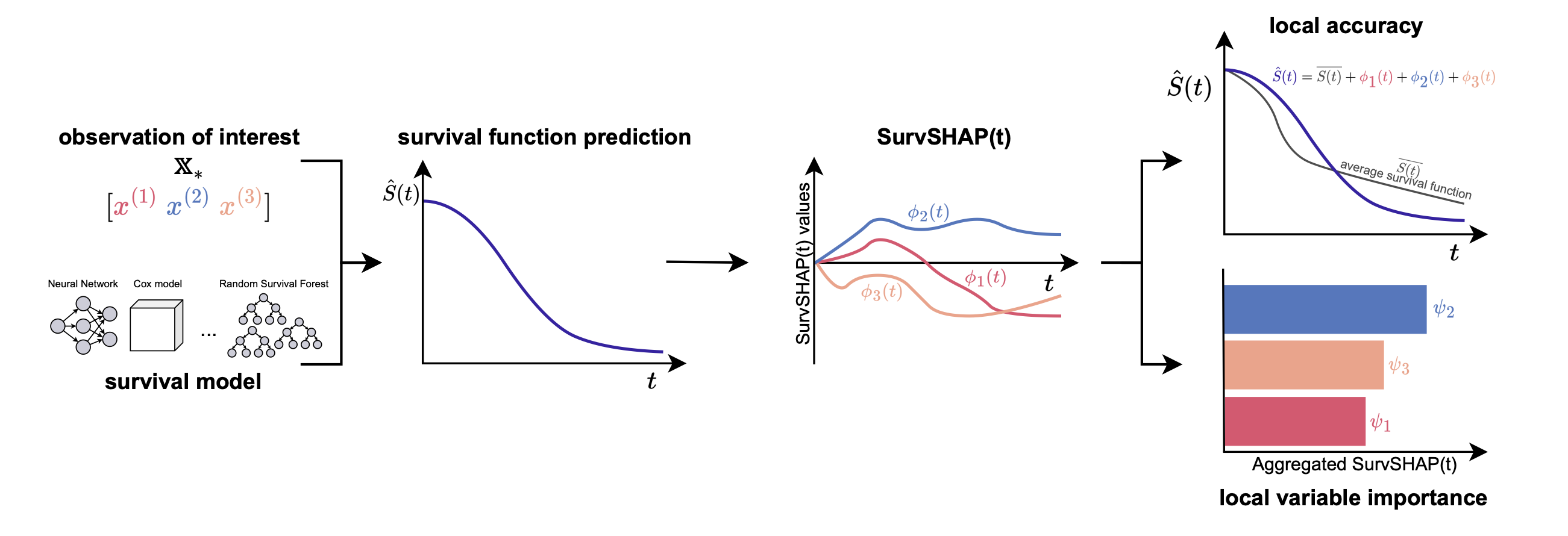
M. Krzyzinski, M. Spytek, H. Baniecki, P. Biecek. SurvSHAP(t): Time-dependent explanations of machine learning survival models. Knowledge-Based Systems 2023
SurvSHAP(t): Shapley values
(Lundberg et al. 2017) SHAP idea: use game theory to estimate additive feature attributions \(\phi_{i}\) to the model’s prediction for observation \(x\).
\[ \phi_{i}(x)=\sum_{S\subseteq P\setminus \{i\}}{\frac {|S|!\;(|P|-|S|-1)!}{|P|!}}(v_{S\cup \{i\}}(x)-v_S(x)) \] \(P\) – feature set, \(v_S(x) = \mathbb{E}_{x|S} f(x)\) – prediction for feature values in set \(S\) that are marginalized over features not included in \(S\).
Over 24 algorithms to estimate Shapley value feature attributions:
KernelSHAP, TreeSHAP, marginal vs conditional..
SurvSHAP(t): example
Idea: explain \(f_t(x)\) – a prediction for an observation \(x\) at time point \(t\) – for all time points separately – \(\phi_t(x, i)\).
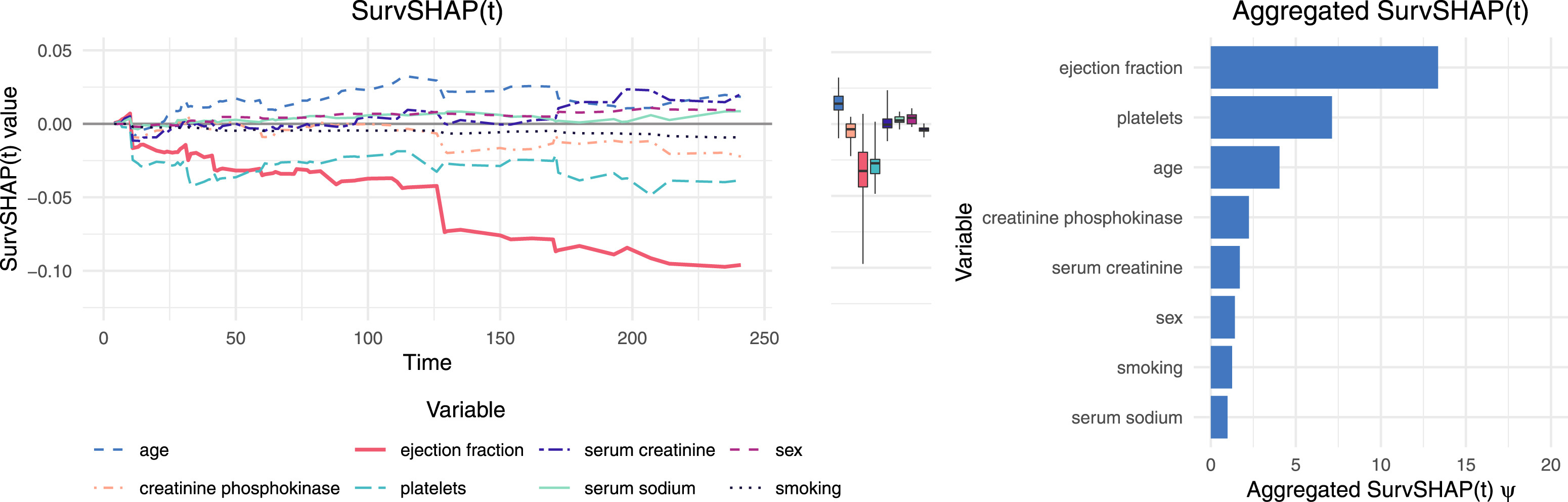
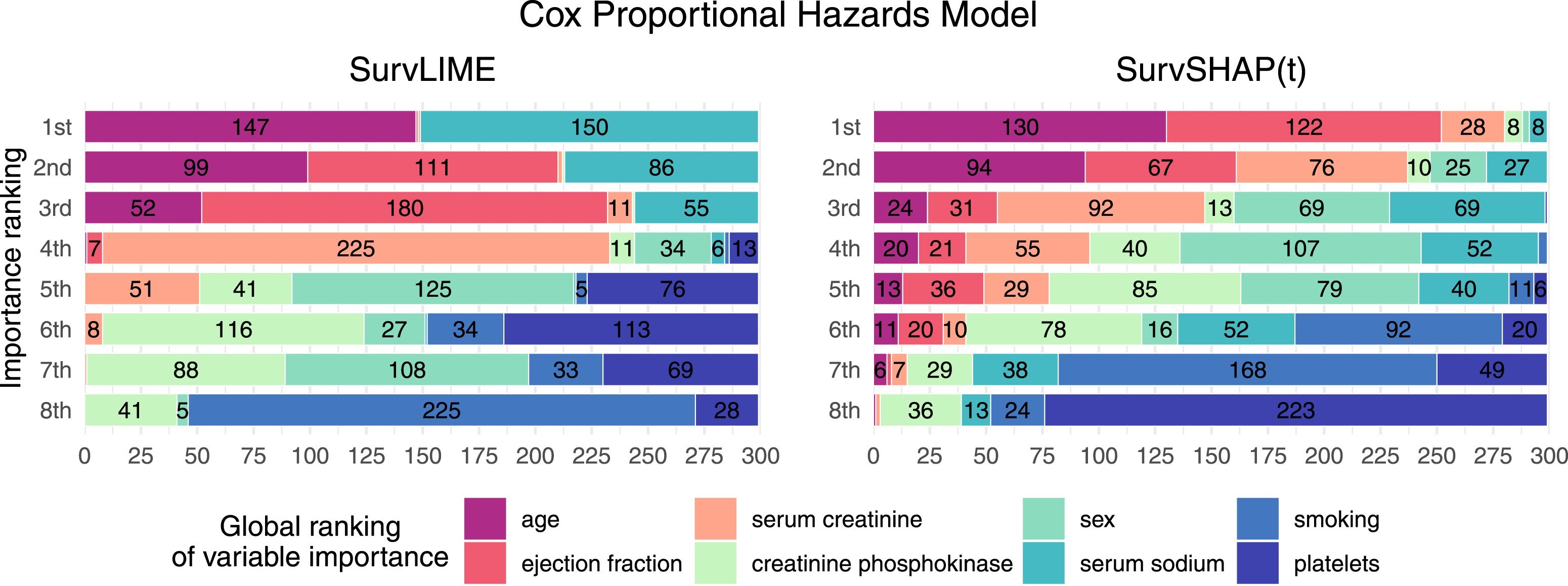
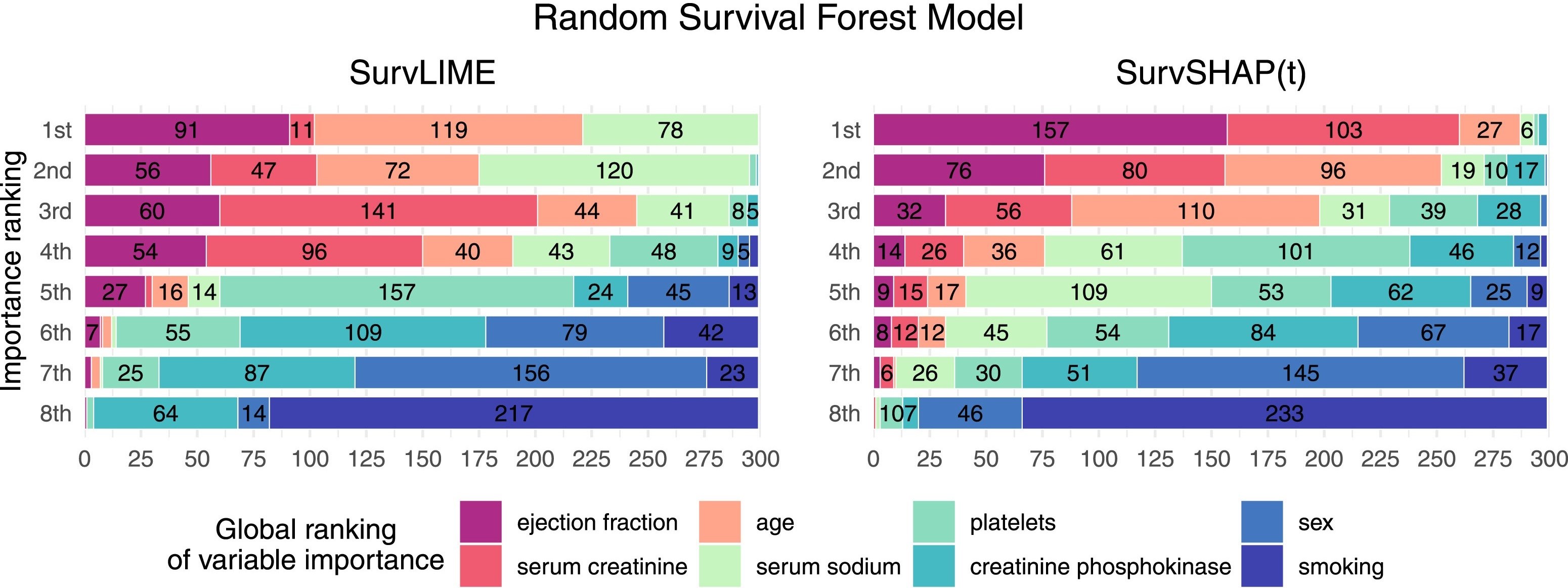
📊 software: survex, survshap
survex: dalex for survival models
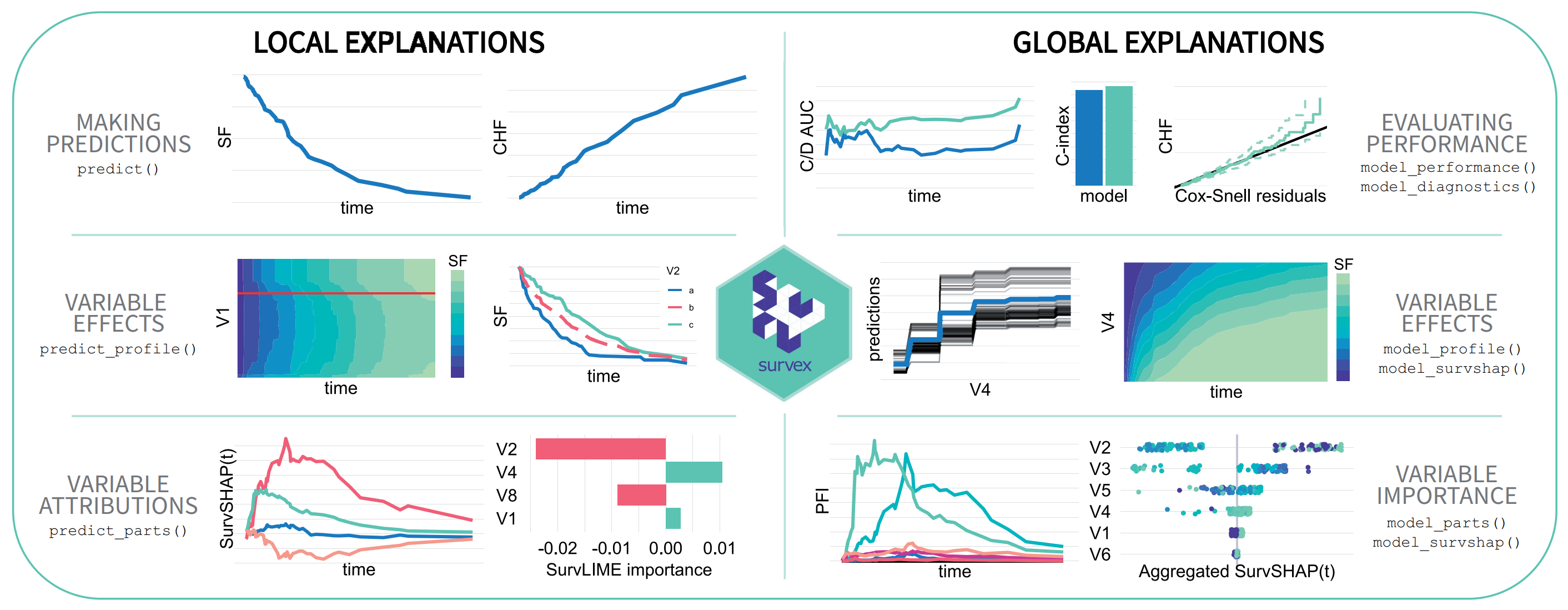
M. Spytek, M. Krzyzinski, S. H. Langbein, H. Baniecki, M. N. Wright, P. Biecek. survex: an R package for explaining machine learning survival models. Bioinformatics 2023
survex: code example
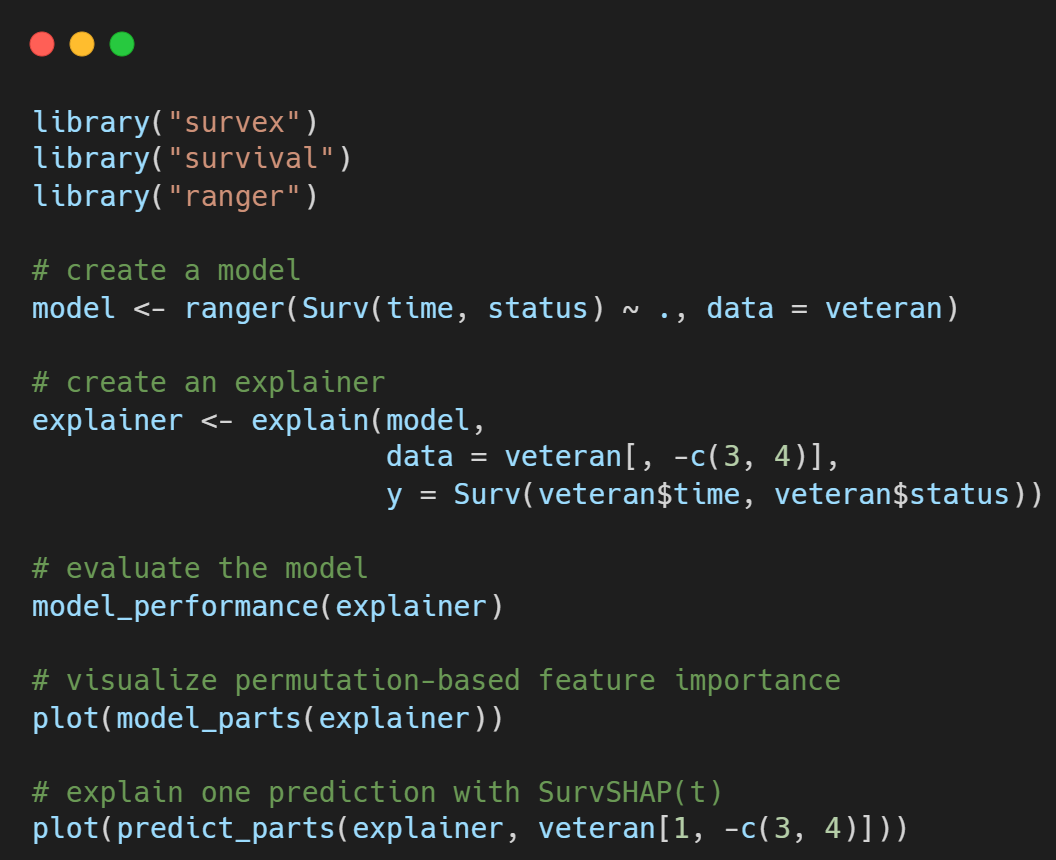
Find more on our GitHub page

🏥 application:
explaining hospital length of stay
Predicting hospital length of stay (LoS)
To what extent can the patient’s length of stay in a hospital be predicted using only an X-ray image?
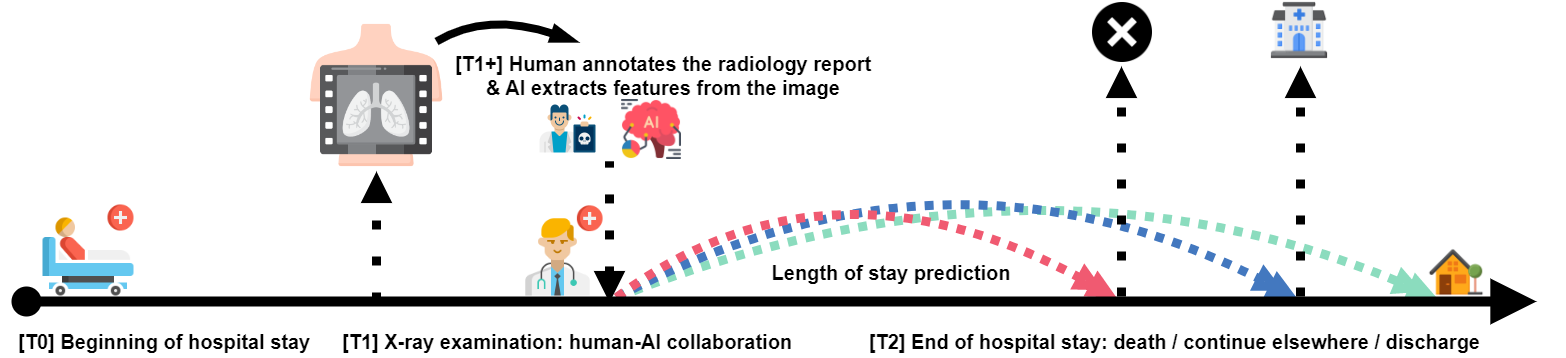
Motivation: explain black-box models predicting hospital LoS
(K. Stone et al. PLOS Digital Health 2022)
Task: time-to-event prediction
instead of a single-value time regression or time-span classification
Dataset i.e. tabular data openly available on GitHub
We manually annotated textual radiology reports of X-ray images:
- 1235 patients from one of the Polish hospitals
- target feature: time between the patient’s radiological examination and hospital discharge (in days, \(min=1\), \(median=7\), \(mean=13.73\), \(max=330\)); about 20% of outcomes are right-censored
- 95 features:
- 2 baseline: age & sex
- 17 human-annotated pathology occurrences, e.g. pulmonary nodule, pleural effusion, medical devices
- 76 algorithm-extracted image statistics using the pyradiomics tool (J. Van Griethuysen et al. Cancer Research 2017)
Benchmarking machine learning models
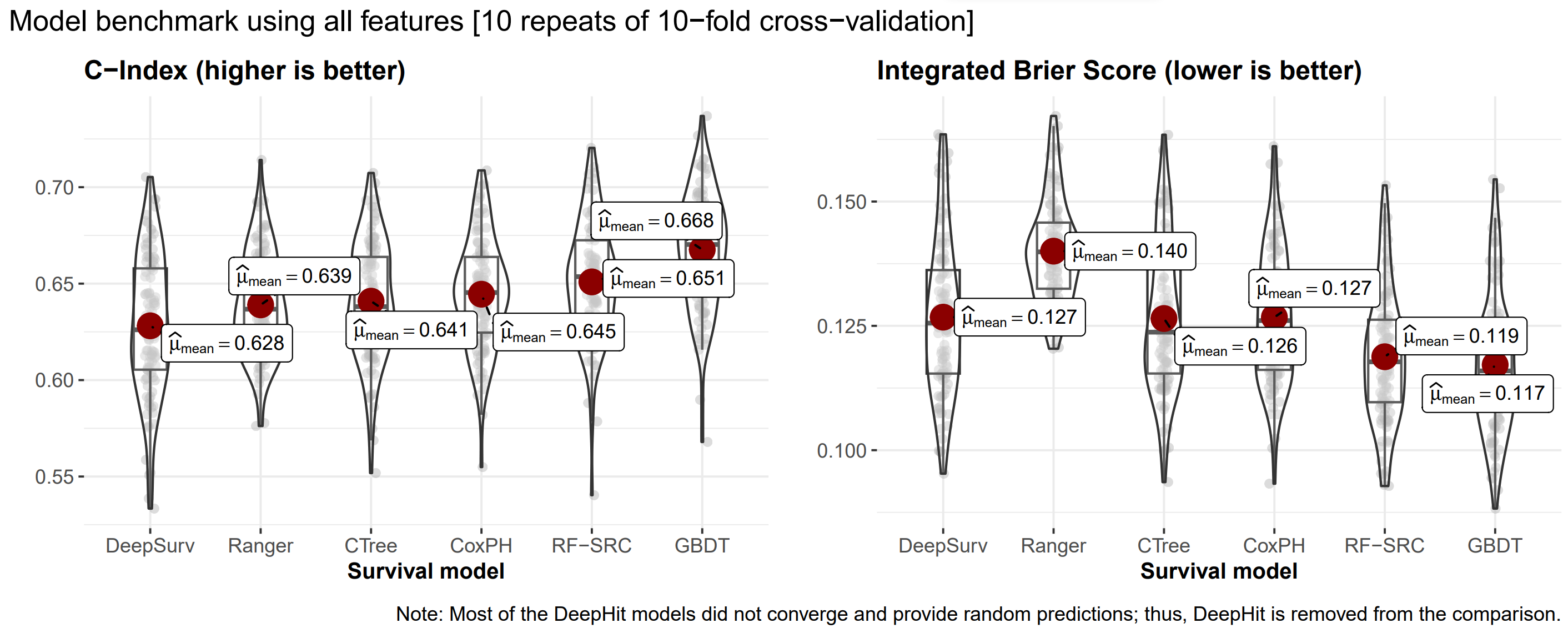
Predicting the patient’s LoS from an X-ray image is indeed possible, but challenging.
Multi-modal feature performance
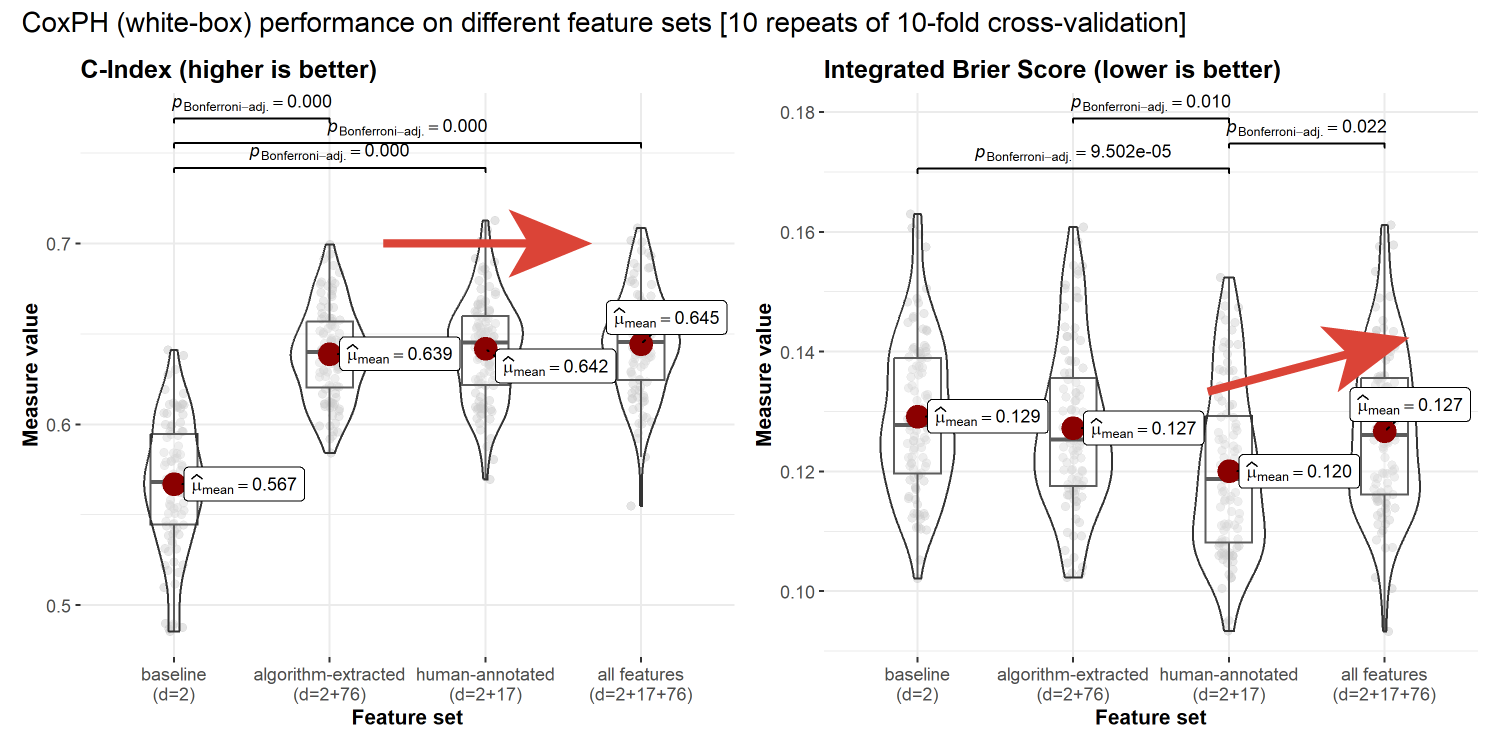
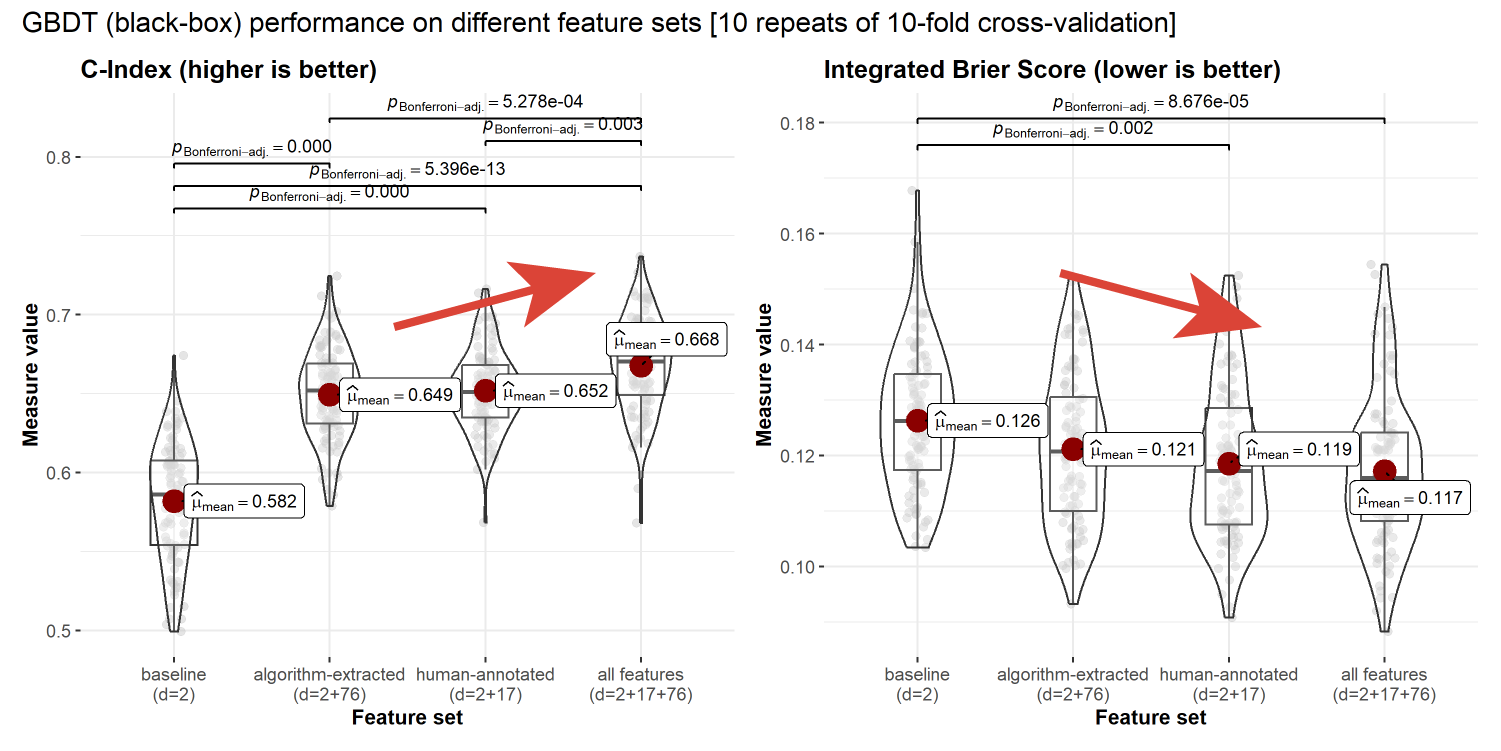
Performance of the interpretable model declines when increasing the number of features.
Explaining LoS predictions to humans
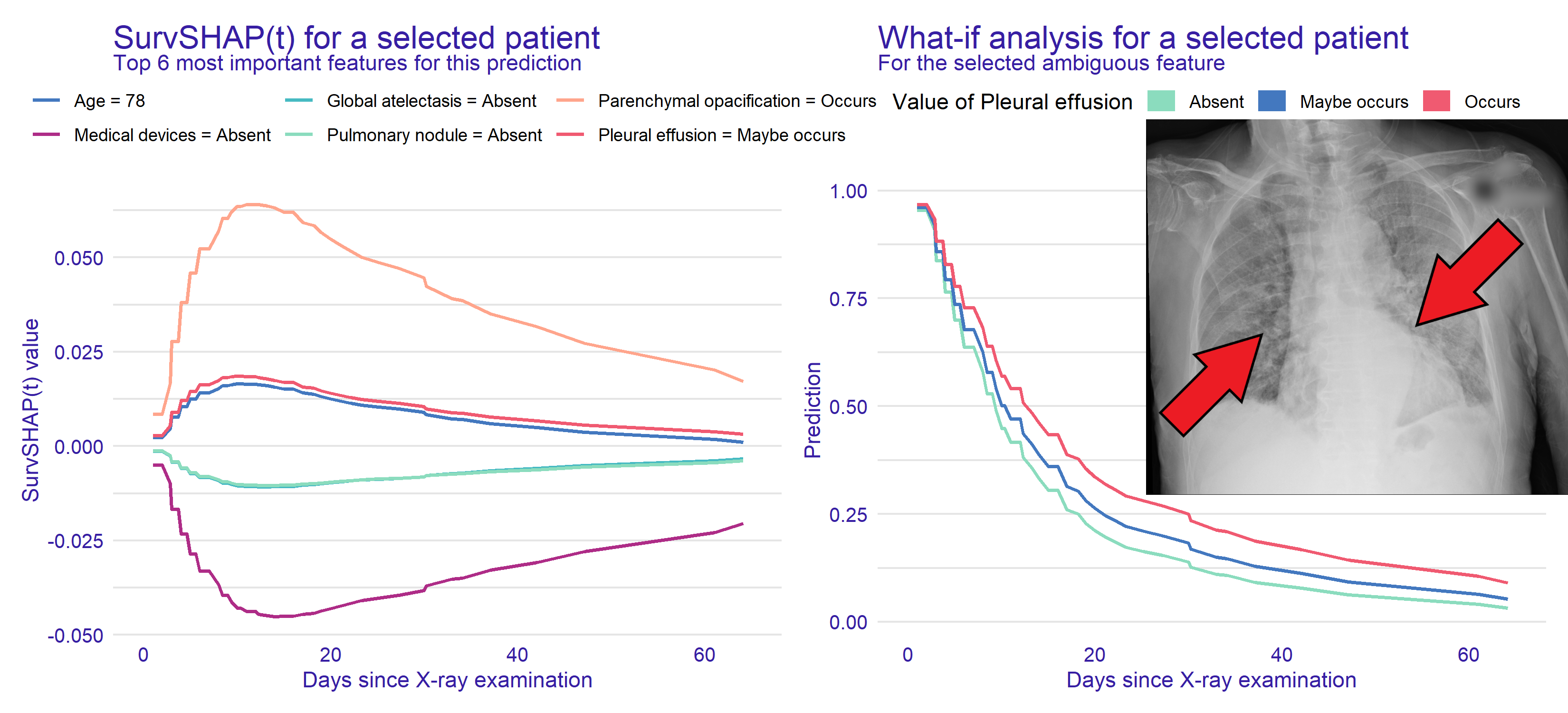
H. Baniecki, B. Sobieski, P. Bombinski, P. Szatkowski, P. Biecek. Hospital Length of Stay Prediction Based on Multi-modal Data Towards Trustworthy Human-AI Collaboration in Radiomics. AIME 2023
Time-dependent feature importance and effects
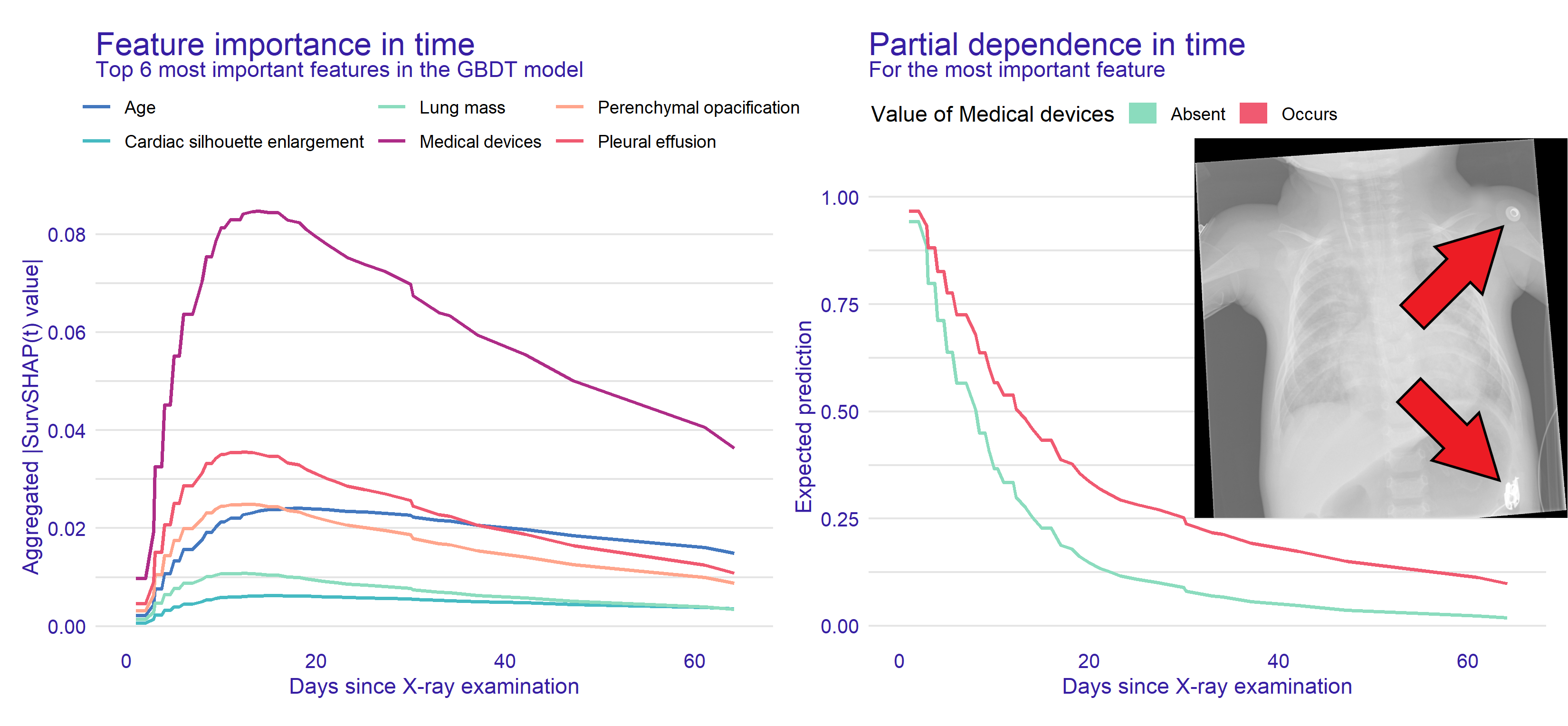
H. Baniecki, B. Sobieski, P. Bombinski, P. Szatkowski, P. Biecek. Hospital Length of Stay Prediction Based on Multi-modal Data Towards Trustworthy Human-AI Collaboration in Radiomics. AIME 2023
Takeaways & discussion
- Adapt explanations to the time-to-event use-case
- The TLOS dataset is openly available for research purposes github.com/mi2datalab/xlungs-trustworthy-los-prediction
How to evaluate explanations❓ How to interpret explanations❓
One needs to be sure that humans properly interpret explanation visualizations, i.e. physicians and other stakeholders interpreting predictions need to understand that explanations are only an approximation of the black-box model.
🧑 🔬 Shameless plug: I’m open to a 3-month research visit in 2025 (self-)funded by “Polish NSF” as part of my PhD.
References
- H.Baniecki, W. Kretowicz, P. Piatyszek, J. Wisniewski, P. Biecek. dalex: Responsible Machine Learning with Interactive Explainability and Fairness in Python. JMLR 2021
- M. Krzyzinski, M. Spytek, H. Baniecki, P. Biecek. SurvSHAP(t): Time-dependent explanations of machine learning survival models. Knowledge-Based Systems 2023
- M. Spytek, M. Krzyzinski, S. H. Langbein, H. Baniecki, M. N. Wright, P. Biecek. survex: an R package for explaining machine learning survival models. Bioinformatics 2023
- H. Baniecki, B. Sobieski, P. Bombinski, P. Szatkowski, P. Biecek. Hospital Length of Stay Prediction Based on Multi-modal Data Towards Trustworthy Human-AI Collaboration in Radiomics. AIME 2023
- K. Stone et al. A systematic review of the prediction of hospital length of stay: Towards a unified framework. PLOS Digital Health 2022
- J. Van Griethuysen et al. Computational radiomics system to decode the radiographic phenotype. Cancer Research 2017

Hubert Baniecki – slides: hbaniecki.com/birs2024 – lab: mi2.ai